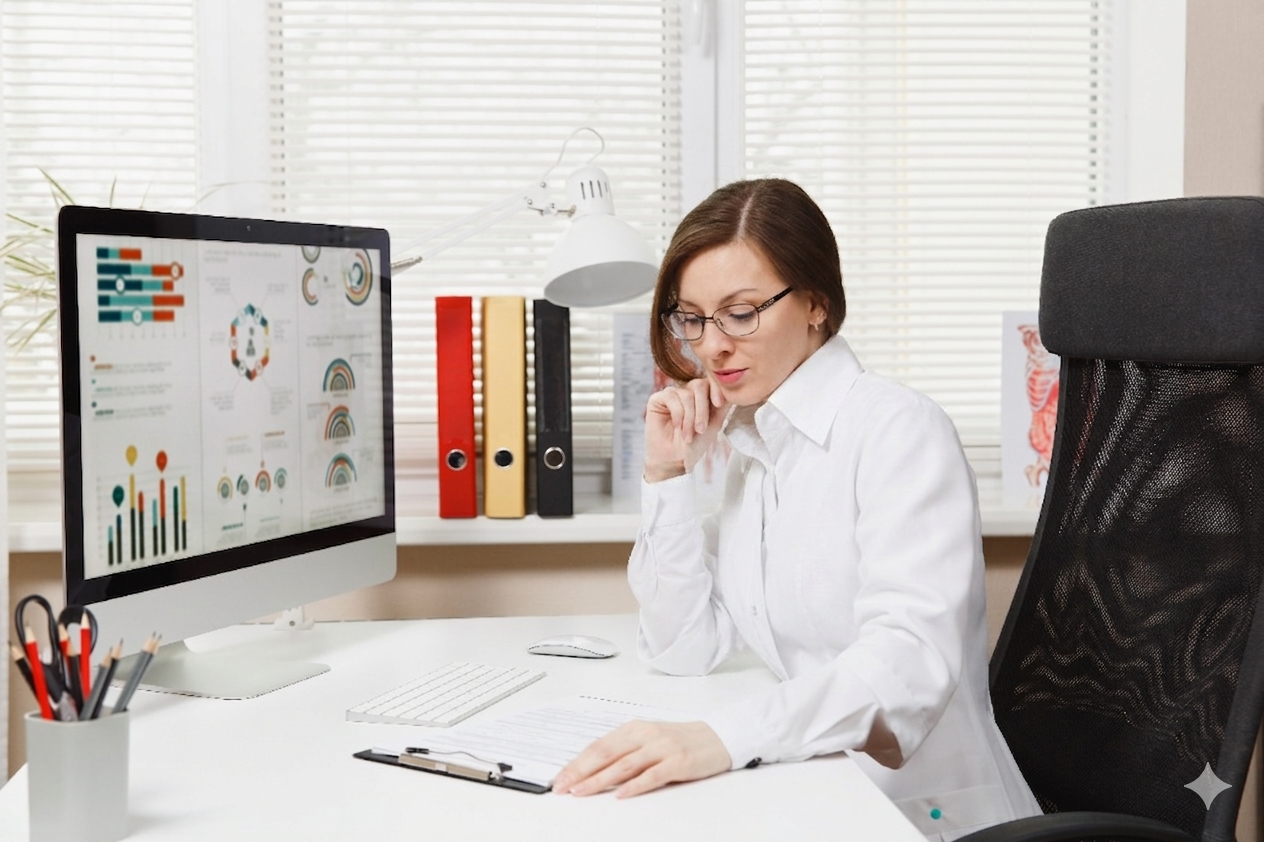
Top Bioinformatics Skills Required in 2026
Bioinformatics skills required in 2026 include NGS analysis, RNA-Seq, variant interpretation, AI-driven biological analysis, cloud computing, multi-omics integration, workflow automation, and biological data interpretation. Employers increasingly value professionals who can combine computational analysis with biological decision-making.
The global bioinformatics market is projected to grow rapidly, reaching approximately USD 32.36 billion in 2025 and expanding to nearly USD 122.89 billion by 2032, with an estimated annual growth rate of about 21%. This sharp expansion reflects how strongly research, healthcare, and biotechnology now depend on computational tools to manage and interpret biological data.
With the rise of high-throughput sequencing, genomics studies, and AI-driven research, the use of advanced bioinformatics software has become essential for handling large-scale biological data. Today, researchers depend on a wide bioinformatics tools list to process, analyze, and interpret complex genomic and molecular datasets efficiently. These tools used in bioinformatics support critical tasks such as sequencing quality checks, read alignment, variant calling, gene expression analysis, protein structure visualization, and reproducible computational workflows.
In real-world research environments, bioinformatics analysis involves multiple interconnected stages, where each tool serves a specific purpose within the data analysis pipeline. For aspiring Bioinformatics Analysts and computational biology professionals, understanding these tools is essential because they form the practical foundation of work in genomics, NGS analysis, structural biology, drug discovery, and precision medicine research. In this blog, we break down the top 13 tools used by Bioinformatics Analysts, organized by domain, so you can clearly understand where each tool fits and why it matters in real-world biological research.
What Are Bioinformatics Tools in Genomics and Drug Discovery?
Bioinformatics tools are specialized computational software used to analyze biological data such as DNA sequences, protein structures, and genomic datasets. These bioinformatics analysis tools help researchers process large datasets generated from sequencing technologies and transform raw biological information into meaningful insights for genomics research, drug discovery, and clinical studies. Modern computational biology tools and genomics analysis tools are widely used for sequence alignment, gene expression analysis, variant detection, protein modeling, and large-scale biological data interpretation.
Who is a Bioinformatics Analyst and What do they do?
A Bioinformatics Analyst works at the intersection of biology and data analysis, transforming complex biological datasets such as DNA sequences, gene expression data, and protein information into meaningful insights for genomics research, drug discovery, disease studies, and clinical research.
Instead of wet-lab experimentation, they use advanced bioinformatics software and computational biology tools to analyze sequencing data, identify patterns, study genetic variations, and interpret large-scale biological information efficiently. These tools play a critical role in modern bioinformatics and genomics workflows.
Top 13 Bioinformatics Tools Used by Bioinformatics Analysts
Bioinformatics analysts rely on specialized bioinformatics tools and bioinformatics software to analyze and interpret complex biological data. These computational biology tools and genomics analysis tools support different stages of research workflows, helping convert raw biological information into meaningful scientific and clinical insights.
A. FASTQC Analysis Tool for NSG Data quality Control
These tools form the foundation of most bioinformatics workflows. They help analysts perform essential tasks such as sequence comparison, genome visualization, quality control, alignment, and statistical analysis. Most beginners start with these tools because they are widely used in genomics and NGS analysis pipelines.
1. BLAST (Basic Local Alignment Search Tool) for sequence similarity & gene identification:
BLAST is one of the most widely used tools for genomics and sequence analysis. It helps bioinformatics analysts compare DNA or protein sequences against large biological databases to identify similar or previously studied sequences. Analysts commonly use BLAST to predict sequence function, identify conserved regions, and study evolutionary relationships across species.
BLAST is a core tool in sequence similarity analysis and is widely integrated into genome annotation and functional analysis workflows. For bioinformatics analysts, understanding BLAST is essential for both sequence comparison and interpreting results alongside downstream tools and scripting platforms like Python or R.
Real‑World Use Case: Identifying Homologous Genes Across Species
In real‑world genomics research, bioinformatics analysts use BLAST to find genes in one species that are like known genes in another. During comparative genomic studies, scientists perform BLAST searches to locate regions of similarity between a query sequence and target genomes. This process helps identify homologous genes, predict their likely function, and understand evolutionary relationships across species.
2. UCSC Genome Browser (genome visualization & variant interpretation):
The UCSC Genome Browser is a widely used genome visualization platform that helps bioinformatics analysts explore chromosomes, genes, mutations, and genomic features through an interactive interface. It is commonly used in human genomics, clinical genomics, population genetics, and disease variant analysis.
Bioinformatics analysts use the UCSC Genome Browser to map genes, examine genomic variants, visualize annotation tracks, and interpret how different genomic elements overlap within specific regions. It is often integrated with NGS pipelines, variant calling tools, and genomics analysis workflows to support mutation interpretation and functional analysis.
Real-World Use Case: Using UCSC Genome Browser in Human Genomics Research
In research studies, analysts used the UCSC Genome Browser to investigate genetic variants in cancer patients. They layered multiple annotation tracks including gene predictions, SNP data, and regulatory elements to pinpoint mutations that might affect gene function. By comparing these regions with data from other species, analysts could also assess evolutionary conservation, helping prioritize variants for further study. This real-world use shows how the browser enables analysts to integrate, visualize, and interpret complex genomic data on one platform.
3. FASTQC – Fast Quality Control for Raw Sequencing Data (Raw Sequencing Data Quality Check):
FASTQC is a widely used quality control tool that evaluates raw sequencing data generated from next-generation sequencing (NGS) experiments. It is commonly used in RNA-seq, DNA-seq, whole-genome sequencing, exome sequencing, and other high-throughput sequencing workflows.
Bioinformatics analysts use FASTQC at the beginning of NGS pipelines to assess sequencing quality through reports on base quality scores, GC content, sequence duplication levels, and other key metrics. This helps identify potential data issues before alignment, variant calling, or downstream analysis. FASTQC is commonly integrated with tools like BWA, HISAT2, and other NGS analysis workflows to ensure reliable sequencing results.
Real‑Life Use Case: Quality Control in RNA‑seq
In RNA‑seq workflows, bioinformatics analysts begin by checking the quality of raw sequencing data to identify low‑quality reads and other issues before further analysis. Analysts use FASTQC to generate quality reports on raw FASTQ files, which highlight metrics such as per‑base quality and GC content, allowing them to trim or filter problematic reads before alignment and downstream processing. This practical step helps improve the accuracy of subsequent read alignment and gene expression analysis.
4. BWA (Burrows–Wheeler-Alignment) / HISAT2 (Hierarchical Indexing for Spliced Alignment of Transcripts 2) – Read Alignment:
BWA and HISAT2 are widely used read alignment tools that map raw sequencing reads from next-generation sequencing (NGS) experiments to a reference genome. They are commonly used in RNA-seq, DNA-seq, whole-genome sequencing, exome sequencing, and transcriptomics workflows. BWA is mainly used for DNA sequencing data, while HISAT2 is optimized for RNA-seq and splice-aware alignment.
Bioinformatics analysts use these tools to accurately place sequencing reads within the genome or transcriptome, forming the foundation for downstream analysis such as variant calling and gene expression studies. BWA and HISAT2 are commonly integrated with FASTQC, SAMtools, and GATK in modern genomics and transcriptomics pipelines.
Real-World Use Case: Read Alignment in RNA-seq
In RNA-seq workflows, bioinformatics analysts use BWA and HISAT2 to map sequencing reads to a reference genome, enabling accurate downstream analyses such as gene expression quantification and variant calling. For instance, a study comparing seven different RNA-seq alignment tools used BWA and HISAT2 on real RNA-seq data from Arabidopsis thaliana to evaluate alignment accuracy and performance, demonstrating their critical role in NGS pipelines.
5. SAMtools – Alignment File Processing (SAM = Sequence Alignment/Map):
SAMtools is a widely used bioinformatics toolkit for processing and managing aligned sequencing reads stored in SAM or BAM file formats. It is commonly applied in RNA-seq, DNA-seq, whole-genome sequencing, and other high-throughput sequencing workflows.
Bioinformatics analysts use SAMtools to sort, index, filter, and manipulate alignment files, ensuring sequencing datasets are properly organized for downstream analysis. In modern NGS pipelines, SAMtools commonly works alongside aligners such as BWA and HISAT2 and downstream tools like GATK, supporting accurate, efficient, and reproducible sequencing analysis workflows.
Real-Life Example: Alignment File Processing in RNA-seq
In a transcriptomics study, analyzing RNA-seq data from human tissue samples, analysts used SAMtools to sort and index BAM files generated by HISAT2 alignment. This allowed them to filter low-quality alignments, efficiently access reads from specific genomic regions, and prepare the dataset for gene expression quantification. Proper file processing using SAMtools ensured that downstream analyses, including differential expression and variant detection, were accurate and reproducible.
6. R (Bioconductor) – Gene Expression & Statistical Analysis:
R is a statistical programming language, and Bioconductor is a collection of R packages designed for high-throughput genomic data analysis. They are widely used in RNA-seq, microarray analysis, differential gene expression studies, functional genomics, and clinical transcriptomics workflows.
Bioinformatics analysts use R and Bioconductor to process genomic datasets, perform statistical testing, analyze gene expression patterns, and generate meaningful visualizations for biological interpretation. In modern sequencing workflows, they commonly integrate with upstream NGS tools such as FASTQC, HISAT2, and SAMtools, supporting reproducible analysis and deeper interpretation of complex biological data.
Real‑World Use Case: End‑to‑End RNA‑seq Differential Expression Analysis
In published genomic research, analysts used R and Bioconductor packages to perform end‑to‑end RNA‑seq analysis starting from preprocessed read counts all the way through differential expression and visualization. This workflow included exploratory analysis of gene expression, statistical testing for differential expression, and visual exploration of results using Bioconductor tools such as DESeq2 and related expression analysis packages. The study provides a comprehensive example of how R/Bioconductor facilitates gene‑level analysis of high‑throughput sequencing data.
7. Galaxy – End-to-End NGS Workflows:
Galaxy is a web-based bioinformatics platform that allows users to perform complete next-generation sequencing (NGS) analysis workflows without extensive command-line coding. It is widely used in RNA-seq, DNA-seq, exome sequencing, metagenomics, and variant calling workflows across research and clinical settings.
Bioinformatics analysts use Galaxy to integrate multiple analysis stages — from quality control and alignment to variant detection and statistical analysis — into reproducible and structured workflows. It commonly works alongside tools like FASTQC, BWA, HISAT2, SAMtools, and GATK, helping researchers manage large-scale sequencing data and build efficient end-to-end bioinformatics pipelines.
Real-World Use Case: Workflow Automation for RNA-seq
In transcriptomics research, analysts used Galaxy to process RNA-seq datasets from human tissue samples. They created a complete workflow including FASTQC for quality control, HISAT2 for read alignment, SAMtools for alignment processing, and feature Counts for gene expression quantification. Using Galaxy allows them to run the entire workflow reproducibly, share it with collaborators, and ensure consistency across multiple datasets.
While basic tools support sequence analysis and NGS workflows, advanced tools focus on structural biology and drug discovery research.
B. Advanced Bioinformatics Tools for Structural Biology and Drug Discovery
These tools are used in advanced research areas such as structural biology, molecular modeling, and computational drug discovery. They allow analysts to study protein structures, simulate molecular interactions, and evaluate potential drug candidates.
8. PyMOL – Protein Structure Visualization:
PyMOL is a molecular visualization tool used to analyze 3D structures of proteins, nucleic acids, and biomolecules at atomic resolution. It helps researchers examine structural geometry, binding pockets, and molecular conformations with precision.
In structural bioinformatics and drug discovery, PyMOL is widely used to visualize protein–ligand interactions, compare structural variants, highlight active sites, and generate publication-quality molecular images. For bioinformatics analysts, it plays a key role in understanding how molecular structure relates to biological function and interaction mechanisms.
Real-World Use Case: Visualizing Protein-Ligand Interactions with PyMOL
In a structural bioinformatics study, researchers investigated potential inhibitors for HIV-1 protease. After predicting binding poses using docking simulations, analysts used PyMOL to visualize the 3D structures of the protein-ligand complexes. By examining hydrogen bonds, hydrophobic interactions, and conformational changes, they could identify key residues involved in binding and select the most promising compounds for further experimental testing. This workflow highlights how PyMOL enables analysts to interpret complex molecular interactions visually, which is critical for structure-based drug discovery.
9. AutoDock – Protein–Ligand Docking:
AutoDock is a computational docking tool used to predict how small molecules (ligands) bind to protein targets. It identifies favorable binding poses and estimates binding energy, helping researchers evaluate the strength and stability of molecular interactions.
In structural bioinformatics and drug discovery, AutoDock is widely used for protein–ligand interaction analysis, virtual screening, and structure-based drug design. Bioinformatics analysts use it to predict binding affinity, prioritize potential drug candidates, and support faster, cost-effective drug discovery workflows before experimental validation.
Real-World Use Case: Protein–Ligand Docking Using AutoDock
In structure-based drug discovery projects, bioinformatics analysts use tools like AutoDock (including AutoDock Vina) to predict how small molecules bind to target proteins. For example, in molecular docking studies of potential inhibitors, analysts run AutoDock Vina to calculate the preferred binding orientations and estimate binding affinities between candidate compounds and protein active sites. This information helps prioritize the most promising molecules for experimental validation and downstream drug development.
10. GROMACS (GROningen MAchine for Chemical Simulations) – Molecular Simulations:
GROMACS is a high-performance molecular dynamics (MD) simulation software used to study the physical movements of atoms and molecules over time. It helps researchers simulate protein folding, protein–ligand interactions, and biomolecular behavior under realistic conditions at atomic resolution.
In bioinformatics and drug discovery, GROMACS is widely used to analyze protein stability, conformational changes, molecular interactions, and binding mechanisms beyond static structural models. Bioinformatics analysts use GROMACS simulations to gain dynamic insights into biomolecular behavior, supporting drug design, protein engineering, and molecular stability analysis before experimental validation.
conformational flexibility and dynamic interactions that static models cannot capture, helping prioritize molecules for further experimental validation.
Recent advances in artificial intelligence are also transforming how researchers predict protein structures and analyze biological dat Real‑World Use Case: Molecular Dynamics Simulation of Protein–Ligand Complex
In computational drug discovery research, bioinformatics analysts use GROMACS to simulate the dynamic behavior of protein–ligand complexes over time. For example, in studies targeting the SARS‑CoV‑2 main protease (3CL‑PRO), researchers performed 100‑nanosecond molecular dynamics simulations using GROMACS to observe how the protein’s structure changes in a solvated environment and to assess stability and interaction patterns of candidate inhibitors. These simulations provide insights into asets.
C. AI-Driven Bioinformatics Tools for Protein Structure Prediction
Artificial intelligence is increasingly transforming bioinformatics research. AI-driven tools can analyze massive biological datasets, predict protein structures, and accelerate drug discovery. These technologies allow researchers to solve complex biological problems faster than traditional computational methods.
11. AlphaFold – AI-Based Structure Prediction:
AlphaFold is an artificial intelligence system that predicts three-dimensional protein structures directly from amino acid sequences using advanced deep learning models. It generates highly accurate structural predictions based on experimentally solved protein structures.
In structural bioinformatics and drug discovery, AlphaFold helps researchers study protein function, identify binding sites, analyze mutations, and accelerate structure-based drug design when experimental methods are unavailable. Bioinformatics analysts use AlphaFold-generated structures in docking studies, molecular dynamics simulations, and functional analysis workflows, supporting faster target validation and therapeutic research.
Real‑World Use Case: Predicting Structures for SARS‑CoV‑2 Proteins
During the COVID‑19 pandemic, researchers used AlphaFold to predict the 3D structures of SARS‑CoV‑2 proteins, including spike proteins across multiple viral variants, when experimental structures were incomplete or unavailable. These high‑accuracy predictions were verified against experimental data and helped researchers analyze structural differences, assess impacts of mutations, and support structure‑based drug screening efforts against the virus
12. GitHub – Version Control & Collaboration:
GitHub is a web-based platform built around Git version control, allowing teams to manage, track, and collaborate on code and computational workflows. It records changes made to scripts, pipelines, and analysis workflows, supporting transparency and reproducibility in research.
In bioinformatics, GitHub is widely used to maintain analysis scripts, manage collaborative projects, document workflows, and support reproducible research practices. Bioinformatics analysts use it to store and share NGS pipelines, statistical analysis scripts, and computational biology projects while ensuring version control, peer review, and workflow consistency.
Real-World Use Case: Collaborative Bioinformatics Workflow Management
In a multi-institution RNA-seq study, bioinformatics analysts used GitHub to manage their workflow scripts, including FASTQC quality checks, HISAT2 alignment, SAMtools processing, and DESeq2 differential expression analysis. By versioning these pipelines on GitHub, all collaborators could access, reproduce, and update the analyses reliably, ensuring consistent results across multiple datasets. This approach improved team productivity, transparency, and reproducibility, making it easier to share and validate workflows in a collaborative research environment.
13. GATK – Variant Calling (GATK = Genome Analysis Toolkit):
GATK (Genome Analysis Toolkit) is a widely used bioinformatics software suite for identifying genetic variants such as SNPs and insertions/deletions (indels) from aligned sequencing data. It is commonly used in whole-genome sequencing, exome sequencing, RNA-seq analysis, clinical genomics, and disease research. Bioinformatics analysts use GATK to process BAM/SAM files and generate high-confidence variant calls through standardized analysis workflows.
Accurate variant detection is critical in modern genomics because downstream applications such as disease studies, population genomics, and precision medicine rely on reliable results. GATK is often integrated with tools like BWA, HISAT2, and SAMtools, making it a core component of end-to-end NGS analysis pipelines.
Real-World Use Case: Variant Calling in Human Genome Studies
In large‑scale sequencing studies such as the 1000 Genomes Project and The Cancer Genome Atlas, bioinformatics analysts use GATK to identify SNPs and other genetic variants from aligned NGS data. After aligning reads with tools like BWA and processing the alignments, GATK’s variant calling functions generate high‑confidence variant calls that help researchers identify mutations linked to traits or diseases.
Bioinformatics
Develop in-depth skills to analyze, manage, and interpret large-scale biological data used in genomics, clinical research, and drug discovery. This program focuses on applying computational methods and analytical pipelines to transform complex biological data into actionable research insights.

Duration: 6 months
Learn at your own pace
Skills you’ll build:
Why Bioinformatics tools are important for a Bioinformatics Career
Bioinformatics tools are the backbone of modern biological research. Large genomic datasets generated from sequencing technologies cannot be analyzed manually, making computational tools essential for processing and interpreting biological information.
Professionals who understand how to use tools such as BLAST, FASTQC, GATK, AlphaFold, and PyMOL can contribute to genomics research, drug discovery, clinical studies, and precision medicine. These tools enable analysts to transform raw biological data into insights that help scientists understand diseases, identify genetic mutations, and develop new therapies.
For aspiring bioinformatics analysts, mastering these tools is not just about learning software it is about understanding how different steps of a bioinformatics workflow connect.
Skills required to master Bioinformatics Analysis tools
To effectively use bioinformatics tools, professionals need a combination of biological knowledge, computational skills, and data analysis expertise.
Key skills include:
• Understanding genomics and molecular biology concepts
• Programming knowledge in Python or R
• Familiarity with Linux command-line environments
• Experience with NGS data analysis workflows
• Statistical analysis and data visualization skills
• Knowledge of databases such as GenBank and PDB
These skills help analysts integrate different tools within a workflow and interpret results accurately.
Why Upskilling and Reskilling are important in Bioinformatics
Bioinformatics is a rapidly evolving field driven by advancements in sequencing technologies, artificial intelligence, and computational biology. As new tools and analysis methods emerge, professionals must continuously upgrade their skills to stay relevant.
Upskilling through structured training programs, workshops, and hands-on projects allows professionals to learn modern bioinformatics workflows, including NGS analysis, structural bioinformatics, and AI-driven drug discovery.
Reskilling is also becoming common for professionals from life sciences, biotechnology, pharmacy, and computer science backgrounds who want to transition into bioinformatics and computational biology roles.
Career Opportunities after mastering Bioinformatics tools
Professionals who develop expertise in bioinformatics tools can pursue careers across biotechnology, pharmaceutical research, healthcare, and academic research.
Common career roles include:
• Bioinformatics Analyst
• Computational Biologist
• Genomics Data Scientist
• Clinical Bioinformatics Specialist
• Drug Discovery Scientist
• NGS Data Analyst
• Research Scientist in Genomics or Proteomics
With the growth of precision medicine and genomic research, the demand for professionals who can analyze biological data using computational tools continues to increase globally.
Conclusion: The Growing Importance of Bioinformatics Tools
Bioinformatics tools transform complex biological datasets into meaningful scientific insights for research and healthcare. From sequence analysis tools like BLAST to AI-driven platforms like AlphaFold, these technologies enable researchers to study genomes, analyze molecular interactions, and accelerate drug discovery.
In this rapidly evolving field, the right combination of practical skills, analytical thinking, and hands-on experience can make all the difference. For those passionate about bridging biology and computation, gaining real-world experience and guidance from industry experts is the key to becoming a confident, job-ready professional.
For those ready to step into this cutting-edge field, practical skills and mentorship are the key. The Advanced Diploma in Bioinformatics at CliniLaunch Research Institute equips learners with hands-on training, industry-standard tools, and real-world projects, preparing you to become a confident, job-ready bioinformatics professional. Start transforming data into discoveries and your career into a breakthrough.
Frequently Asked Questions (FAQs)
NGS analysis commonly uses FastQC for quality control, BWA and Bowtie for sequence alignment, SAMtools for data processing, and GATK for variant calling. These tools help researchers analyze large-scale sequencing data efficiently.
Some beginner-friendly bioinformatics software includes BLAST, Galaxy, MEGA, PyMOL, and Bioconductor. These tools are widely used for sequence analysis, visualization, and genomics research while offering accessible learning environments for beginners.
Drug discovery workflows often use bioinformatics tools such as AutoDock, PyMOL, AlphaFold, SwissADME, and molecular modeling platforms for protein structure analysis, target identification, and compound interaction studies.
Python is highly valuable in bioinformatics because it helps automate workflows, analyze biological datasets, and handle large-scale genomic data. Many bioinformatics analysts use Python alongside bioinformatics software for data processing and computational analysis.
Computational biology tools include BLAST, Clustal Omega, Bioconductor, GROMACS, Cytoscape, and R programming environments. These tools support genomics analysis, protein modeling, biological network analysis, and large-scale data interpretation.
Several widely used bioinformatics tools are open source, including Galaxy, Bioconductor, SAMtools, FastQC, BWA, and Bowtie. These tools are popular in research environments because they are freely accessible and supported by active scientific communities.