A biology degree today extends far beyond traditional laboratory research, academia, or teaching. The modern biosciences landscape has evolved into a broad, industry-driven ecosystem where biological knowledge supports real-world healthcare delivery, clinical trials, regulatory systems, digital health platforms, and scientific communication. As healthcare becomes more structured and technology-enabled, biology graduates are increasingly contributing in applied, operational, and analytical roles.
The life sciences industry now functions through interconnected domains such as clinical research and drug development, healthcare data analytics and AI integration, regulatory compliance and quality systems, medical documentation, and clinical operations management. These sectors rely not only on experimentation, but also on documentation accuracy, patient safety oversight, data interpretation, and coordinated execution across multidisciplinary teams.
With continued expansion in pharmaceuticals, biologics, vaccines, and digital health technologies, structured non-laboratory roles are steadily increasing. For biology graduates, the opportunity lies in identifying where industry demand exists and strategically aligning their scientific foundation with practical, role-specific skills that match current hiring patterns.
The Biology Advantage: Transferable Industry Skills
Biology graduates possess foundational strengths that translate directly into life sciences industries:
- Understanding of human physiology and disease mechanisms
- Familiarity with research methodologies
- Medical terminology comprehension
- Awareness of ethics in human research
- Analytical interpretation of data
- Structured documentation exposure
- Scientific reading and comprehension ability
These foundational competencies reduce the learning curve in industry roles where biological context matters.
High-Value Skill Combinations That Increase Employability
Modern hiring favors hybrid skill profiles. Biology alone is foundational, but biology combined with applied industry skills significantly improves opportunities.
High-demand combinations include:
- Biology + Clinical Research
- Biology + Data Analytics
- Biology + AI / Machine Learning
- Biology + Regulatory Compliance
- Biology + Scientific Writing
- Biology + Project Management
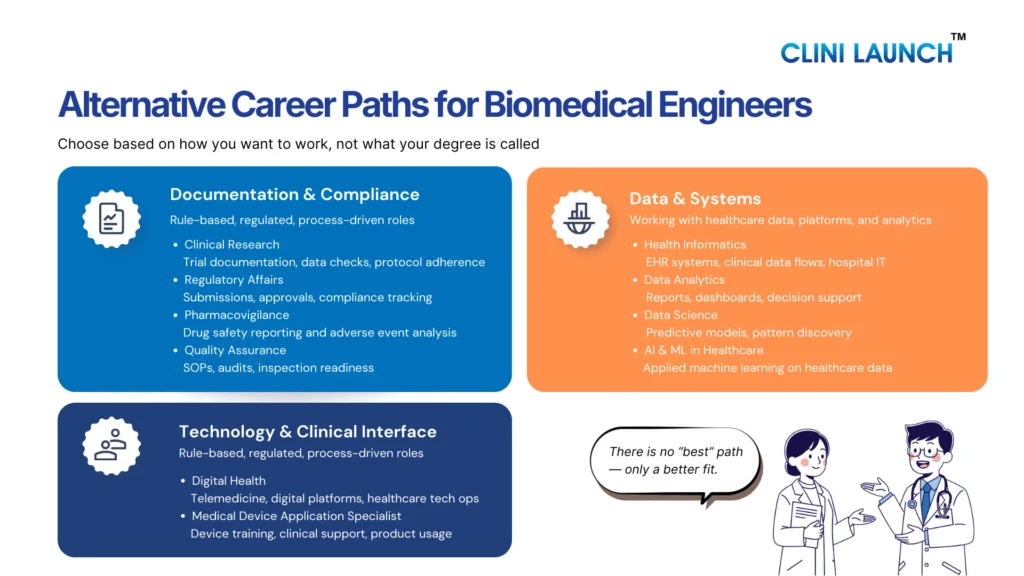
Alternative Careers for Biosciences
Choosing the right career path becomes easier when you understand what you naturally enjoy. If you like working with numbers, logic, or coding, careers in bioinformatics or healthcare data analytics may suit you. If you are detail-oriented and comfortable following rules and documentation processes, regulatory affairs or medical coding can be good options.
If you enjoy reading, writing, and explaining scientific topics, medical writing or pharmacovigilance may be a better fit. If you are curious about how technology is used in healthcare, areas like health informatics or AI in healthcare are growing fields to explore. Identifying your interests helps you choose a direction where your biology background can be applied confidently and practically.
A. Clinical Research & Drug Development
The Clinical Research & Drug Development sector focuses on how new drugs, vaccines, biologics, and medical devices are tested in human subjects before regulatory approval. Clinical trials are conducted in structured phases (Phase I–IV) to evaluate safety, efficacy, dosage, and long-term outcomes.
This industry follows strict global guidelines such as ICH-GCP to protect patient safety, ensure reliable and ethical research practices, and maintain accurate documentation for regulatory compliance and audits.
Clinical research involves collaboration between sponsors (pharma companies), CROs (Contract Research Organizations), hospitals, ethics committees, and regulatory authorities.
1. Clinical Trial Assistant (CTA)
Clinical Trial Assistants support the administrative and documentation of backbone clinical trials. They work closely with Clinical Research Associates and project teams to maintain trial master files (TMF), track essential documents, coordinate communications with trial sites, and ensure that study records remain inspection ready. The role is process-driven and office-based, focusing on compliance, documentation accuracy, and operational coordination rather than laboratory work.
Specific Skills Required:
- Understanding of ICH-GCP guidelines
- Clinical trial documentation management (TMF handling)
- Basic knowledge of study protocols and visit schedules
- Regulatory document tracking
- Strong organizational and communication skills
- Familiarity with eTMF systems (in growing organizations)
Entry Salary (India):
₹3–5 LPA for freshers, depending on location, employer type (CRO, sponsor, hospital), and practical exposure.
Growth Areas:
- Risk-Based Monitoring (RBM) models
- Electronic Trial Master File (eTMF) systems
- Decentralized Clinical Trials (DCTs)
- Global multi-country study coordination
2. Clinical Research Associate (CRA)
Clinical Research Associates are responsible for monitoring clinical trial sites to ensure that studies are conducted according to approved protocols, ICH-GCP guidelines, and regulatory requirements. CRAs act as the link between sponsors (pharma companies or CROs) and investigative sites (hospitals or research centers). Their primary responsibility is to verify that patient safety is protected, trial data is accurate, and documentation is compliant and audit ready. The role involves site visits (on-site or remote), source data verification (SDV), investigator communication, and issue resolution during the trial lifecycle.
Specific Skills Required:
- Strong understanding of ICH-GCP and regulatory frameworks
- Clinical trial monitoring procedures
- Source data verification (SDV)
- Protocol interpretation and deviation handling
- Site management and stakeholder communication
- Risk identification and documentation review
- Report writing (monitoring visit reports)
Entry Salary (India):
For entry-level or junior CRA roles (often after CTA/CRC experience), salaries typically range from ₹4–6 LPA, depending on organization type, location, and prior exposure. With 2–4 years of experience, compensation increases significantly.
Growth Areas:
- Risk-Based Monitoring (RBM)
- Remote and centralized monitoring models
- Global multi-country trials
- Oncology and biologics trials
- Digital clinical trial platforms

3.Clinical Research Coordinator (CRC)
Clinical Research Coordinators work at the trial site level (hospitals, research centers, or investigator sites) and are responsible for the day-to-day coordination of clinical studies. They ensure that trials are conducted according to approved protocols, ethical guidelines, and regulatory requirements. CRCs act as the central point of contact between investigators, patients, sponsors, and CROs. Their responsibilities include patient screening and enrollment, coordinating study visits, maintaining site documentation, and supporting compliance during audits and monitoring visits. The role is patient-facing and operational rather than laboratory-based.
Specific Skills Required:
- Understanding of ICH-GCP and clinical trial processes
- Patient recruitment and informed consent handling
- Study visits coordination and scheduling
- Source document maintenance
- Ethics committee submission support
- Data entry and query resolution
- Communication and coordination skills
Entry Salary (India):
Typically ranges between ₹3–5 LPA, depending on hospital type, research unit size, city, and prior training exposure.
Growth Areas:
- Multi-specialty hospital research units
- Oncology and specialty trials
- Site management organizations (SMOs)
- Transition into CRA or Site Manager roles
4.Clinical Research Coordinator (CRC)
Clinical Research Coordinators work at the trial site level (hospitals, research centers, or investigator sites) and are responsible for the day-to-day coordination of clinical studies. They ensure that trials are conducted according to approved protocols, ethical guidelines, and regulatory requirements. CRCs act as the central point of contact between investigators, patients, sponsors, and CROs. Their responsibilities include patient screening and enrollment, coordinating study visits, maintaining site documentation, and supporting compliance during audits and monitoring visits. The role is patient-facing and operational rather than laboratory-based.
Specific Skills Required:
- Understanding of ICH-GCP and clinical trial processes
- Patient recruitment and informed consent handling
- Study visits coordination and scheduling
- Source document maintenance
- Ethics committee submission support
- Data entry and query resolution
- Communication and coordination skills
Entry Salary (India):
Typically ranges between ₹3–5 LPA, depending on hospital type, research unit size, city, and prior training exposure.
Growth Areas:
- Multi-specialty hospital research units
- Oncology and specialty trials
- Site management organizations (SMOs)
- Transition into CRA or Site Manager roles
5.Pharmacovigilance / Drug Safety Associate
Pharmacovigilance (PV) or Drug Safety Associates are responsible for monitoring, assessing, and reporting adverse drug reactions (ADRs) and other safety-related information associated with pharmaceutical products. Their primary role is to ensure patient safety after a drug enters clinical trials or the market. They review safety reports, evaluate case data, perform medical coding of adverse events, and submit reports to regulatory authorities within defined timelines. This role is largely documentation-driven and analytical, requiring careful evaluation of clinical information rather than laboratory work.
Specific Skills Required:
- Understanding of drug safety regulations and reporting timelines
- Knowledge of ICH guidelines related to safety reporting
- Adverse event case processing
- Medical coding using tools such as MedDRA and WHO-Drug dictionaries
- Narrative writing for safety reports
- Data review and signal detection basics
- Attention to detail and regulatory compliance awareness
Entry Salary (India):
Typically ranges between ₹3–5 LPA for freshers, depending on employer type (CRO, pharma company, KPO), location, and safety database exposure.
Growth Areas:
- Signal detection and risk management
- Aggregate safety reporting (PSUR, DSUR)
- Risk Management Plans (RMP)
- Global safety database systems
- Transition into Drug Safety Specialist or PV Manager roles
Clinical Research
Gain in-depth understanding of how clinical trials operate across pharma companies, CROs, and research sites. This program focuses on the practical execution of clinical studies, ethical conduct, documentation, and regulatory compliance across the clinical trial lifecycle.
B. Healthcare Data & Bioinformatics
The Healthcare Data & Bioinformatics sector focuses on converting medical and biological data into insights that improve patient care, drug development, and healthcare decisions. With widespread digitization, data from electronic health records, clinical trials, genomics, imaging, and real-world evidence has become central to modern healthcare operations.
Bioinformatics uses computational and statistical tools to analyze biological data, especially in genomics and molecular biology, supporting disease research and precision medicine. Healthcare data analytics focuses on interpreting clinical and operational datasets to enhance treatment strategies, regulatory reporting, and healthcare efficiency.
With the rise of AI and machine learning, the field now enables predictive modeling for disease risk, drug discovery, and clinical trial optimization. As digital healthcare expands, professionals who combine biological knowledge with data and technology skills are increasingly in demand.
1.Healthcare Data Analyst
Healthcare Data Analysts work with clinical, operational, and patient datasets to extract meaningful insights that support medical decisions, healthcare planning, and research outcomes. They analyze structured data from electronic health records (EHRs), clinical trials, insurance claims, and hospital systems to identify trends, measure treatment effectiveness, and improve care delivery. The role bridges healthcare knowledge with data analysis, focusing on interpretation rather than pure programming.
Specific Skills Required:
- Strong foundation in statistics and data interpretation
- Proficiency in Excel and SQL
- Basic knowledge of Python or R for data analysis
- Understanding of healthcare datasets and medical terminology
- Data visualization tools (Power BI, Tableau, or similar)
- Analytical thinking and attention to accuracy
Entry Salary (India):
Typically ranges between ₹4–7 LPA, depending on technical skill level, tool proficiency, and employer type (health-tech firm, pharma analytics unit, hospital system, or CRO).
Growth Areas:
- Real-World Evidence (RWE) analytics
- Predictive healthcare modeling
- AI-assisted clinical analytics
- Population health analytics
- Transition into Healthcare Data Scientist roles
2.Bioinformatics Analyst
Bioinformatics Analysts use computational tools and statistical methods to analyze biological data, particularly genomic, transcriptomic, and proteomic datasets. They work on sequence analysis, gene expression studies, variant identification, and biomarker discovery to support research, drug development, and precision medicine initiatives. The role sits at the intersection of biology, computer science, and statistics, requiring both domain understanding and technical capability.
Specific Skills Required:
- Strong foundation in molecular biology and genetics
- Knowledge of sequence alignment and genomic databases (e.g., NCBI, Ensembl)
- Familiarity with tools such as BLAST and genome analysis pipelines
- Basic programming skills (Python, R, or similar)
- Statistical analysis and data interpretation
- Understanding of next-generation sequencing (NGS) data
Entry Salary (India):
Typically ranges between ₹4–6 LPA for entry-level roles, depending on programming skills, research exposure, and employer type (biotech firms, research labs, pharma R&D units).
Growth Areas:
- Genomic data analysis and NGS platforms
- Precision medicine and biomarker research
- AI-driven drug discovery
- Computational biology in biotech startups
- Transition into Computational Biologist or Bioinformatics Scientist roles
Bioinformatics
Build practical skills to analyze and interpret biological data generated from genomics, proteomics, and clinical research studies. Learn how computational tools and data-driven methods are used to convert raw biological data into meaningful insights for research and drug development.
C. Regulatory Affairs & Quality Systems
The Regulatory Affairs & Quality Systems sector ensures that pharmaceutical products, biologics, medical devices, and clinical trials meet national and international regulatory standards before and after market approval. Regulatory teams prepare and submit structured dossiers containing clinical, safety, manufacturing, and labeling data to authorities such as CDSCO, US FDA, and EMA to obtain and maintain product approvals.
Quality systems operate alongside regulatory functions to ensure continuous compliance with Good Practice standards such as GMP, GLP, and GCP. This includes developing SOPs, conducting audits, managing deviations and CAPA processes, and maintaining readiness inspections. Compliance is mandatory for companies to manufacture, market, and export healthcare products.
As global regulations become stricter and more harmonized, regulatory and quality roles have evolved into strategic functions within pharmaceutical and biotech organizations. In India, these roles remain stable and essential due to expanding drug exports, biosimilars development, global clinical trial participation, and increasing regulatory scrutiny.
1.Regulatory Affairs Associate
Regulatory Affairs Associates are responsible for preparing, compiling, and submitting documentation required for regulatory approval of pharmaceutical products, biologics, or medical devices. They ensure that products meet national and international regulatory requirements throughout development, approval, and post-marketing stages. The role involves reviewing clinical data, manufacturing information, labeling details, and ensuring submissions comply with guidelines issued by authorities such as CDSCO (India), US FDA, EMA, and other global regulators. It is documentation-intensive and requires strong regulatory understanding rather than laboratory work.
Specific Skills Required:
- Understanding of regulatory frameworks (India and global markets)
- Knowledge of dossier formats such as CTD/eCTD
- Familiarity with clinical and manufacturing documentation
- Regulatory submission process awareness
- Attention to detail and documentation accuracy
- Ability to interpret guidelines and regulatory updates
- Strong written and communication skills
Entry Salary (India):
Typically ranges between ₹3–5 LPA for entry-level roles, depending on company size, export exposure, and regulatory market focus.
Growth Areas:
- Global regulatory submissions (US, EU, emerging markets)
- Biologics and biosimilars regulation
- Regulatory intelligence and strategy
- eCTD publishing and digital submissions
- Transition into Regulatory Manager or Regulatory Lead roles
overview:
AI Integration in Drug Safety and Compliance
Learn how artificial intelligence is applied in pharmacovigilance, drug safety operations, and regulatory compliance. This program focuses on automating safety workflows, improving signal detection, enhancing case processing, and strengthening global regulatory reporting using AI-driven systems.
2.Quality Assurance (QA) Executive
Quality Assurance Executives ensure that pharmaceutical, biotechnology, or medical device operations comply with established quality standards and regulatory requirements. Their role focuses on maintaining Good Practice standards such as GMP (Good Manufacturing Practice), GCP (Good Clinical Practice), or GLP (Good Laboratory Practice), depending on the organization. They monitor processes, review documentation, handle deviations, support audits, and ensure that systems are compliant and inspection ready. QA is process-driven and compliance-focused rather than research-based.
Specific Skills Required:
- Understanding of GMP, GCP, or GLP guidelines
- SOP development and review
- Deviation handling and CAPA (Corrective and Preventive Action) management
- Audit preparation and documentation review
- Risk assessment and quality documentation control
- Attention to detail and regulatory awareness
Entry Salary (India):
Typically ranges between ₹3–5 LPA for freshers, depending on industry segment (manufacturing, clinical research, biotech) and organization size.
Growth Areas:
- Quality Management Systems (QMS) digitization
- Data integrity compliance
- Audit and inspection leadership
- Validation and risk management roles
- Progression to QA Manager or Quality Head positions
D. Medical Writing & Scientific Communication
This sector focuses on converting complex clinical and scientific data into clear, accurate, and regulatory-compliant documents. These documents support drug development, clinical trials, regulatory submissions, and medical education.
Clinical documentation includes study protocols, clinical study reports, investigator brochures, safety narratives, and informed consent forms. Regulatory writing ensures that trial data is presented in standardized formats required by authorities for product approval.
As clinical research expands and regulatory scrutiny increases, the need for professionals who can interpret scientific data and communicate it clearly remains strong across pharmaceutical companies, CROs, and medical communication agencies.
1.Medical Writer
Medical Writers develop structured scientific and clinical documents that support drug development, regulatory submissions, and medical communication. They work with clinical trial data, safety information, and research findings to prepare clear, accurate, and guideline-compliant documents such as clinical study reports (CSRs), protocols, investigator brochures, and safety narratives. The role requires strong scientific understanding combined with precise and structured writing skills.
Specific Skills Required:
- Strong comprehension of clinical research and medical terminology
- Ability to interpret clinical trial data
- Structured scientific writing skills
- Familiarity with ICH guidelines and regulatory document formats
- Literature review and referencing skills
- Attention to detail and consistency
Entry Salary (India):
Typically ranges between ₹3–5 LPA for entry-level positions, depending on writing proficiency, documentation exposure, and organization type (CRO, pharma company, medical communication agency).
Growth Areas:
- Regulatory writing specialization
- Aggregate safety report writing (PSUR, DSUR)
- Manuscript and publication writing
- Medical education and training content
- Progression to Senior Medical Writer or Lead Writer roles
2. Scientific Content Developer
Scientific Content Developers create accurate, evidence-based educational and medical materials for healthcare professionals, pharmaceutical companies, training platforms, and digital health organizations. Their work involves interpreting scientific research and presenting it in simplified, audience-appropriate formats such as learning modules, medical education materials, website content, product explainers, slide decks, and healthcare awareness resources. Unlike regulatory writing, this role focuses more on knowledge of translation and structured communication rather than formal submission documents.
Specific Skills Required:
- Strong understanding of life sciences and medical concepts
- Scientific literature review and interpretation
- Ability to simplify complex information clearly
- Structured writing and content organization
- Basic referencing and citation skills
- Familiarity with medical communication standards
Entry Salary (India):
Typically ranges between ₹3–5 LPA for entry-level roles, depending on writing quality, subject knowledge, and employer type (ed-tech, medical communication agency, pharma marketing team, digital health platform).
Growth Areas:
- Digital medical education platforms
- Healthcare e-learning development
- Pharma product training content
- Scientific communication strategy
- Progression to Senior Content Strategist or Medical Communication Manager
Conclusion
Choosing an alternative career path does not mean moving away from biology. It means applying biological knowledge in areas where today’s life sciences and healthcare industries operate. As clinical research, data-driven healthcare, regulatory systems, and digital workflows continue to grow; these roles are becoming essential to how scientific knowledge is translated into real-world outcomes.
For biology majors, the key is to move beyond uncertainty and focus on building practical, industry-aligned skills. This is where structured learning and guided exposure make a real difference. At CliniLaunch Research Institute, programs are designed to help biology graduates understand industry workflows, gain hands-on exposure, and prepare for entry-level roles with clarity and confidence. With the right direction and preparation, alternative careers offer stable growth, meaningful work, and a future where biology remains a strong foundation for long-term success.
FAQ
1. What jobs are similar to biologist?
Roles such as Clinical Research Associate, Bioinformatics Analyst, Regulatory Affairs Associate, Medical Writer, and Healthcare Data Analyst use biological knowledge in applied industry settings beyond laboratory work.
2. How to transition out of biology?
Add industry-relevant skills like clinical research training, data analytics tools, regulatory documentation, or medical writing. Short-term certifications help shift from academic biology to applied roles.
3. What else can I do with a biology degree?
You can enter clinical research, pharmacovigilance, regulatory affairs, healthcare analytics, bioinformatics, medical writing, or quality assurance without staying in pure lab research.
4. What jobs can I get with just biology?
Entry-level roles include Clinical Trial Assistant, Clinical Research Coordinator, QA Executive, Pharmacovigilance Associate, or Medical Coding Executive. Skill enhancement improves growth opportunities.
5. Which field will boom in 2025?
Healthcare data analytics, AI in healthcare, bioinformatics, and digital clinical trials are expected to grow due to increasing healthcare digitization and global drug development expansion.
6. What is the best career option for biology students?
The best option depends on your interests—data roles suit analytical minds, regulatory suits detail-oriented individuals, and clinical research suits those interested in patient-facing environments.
7. What job should I get if I like biology?
Consider clinical research, drug safety, bioinformatics, regulatory affairs, or healthcare analytics—these fields apply biology in structured industry roles.
8. What is the highest paid biology job?
Healthcare Data Scientist, Bioinformatics Scientist, Clinical Project Manager, and Regulatory Affairs Manager are among the higher-paying biology-related careers with experience.
9. Which branch of biology is most in demand?
Bioinformatics, clinical research, regulatory sciences, and genomics are currently in strong demand due to personalized medicine and regulatory expansion.
10. How useful is a bachelor’s in biology?
A biology degree provides a strong scientific foundation and becomes highly valuable when combined with applied industry skills and practical training.